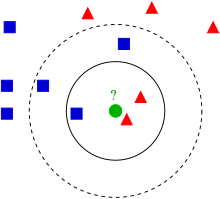
使用KNN算法进行分类。
Instructions
- Do not import other libraries. You are only allowed to use Math, Numpy packages which are already imported in the file. DO NOT use scipy functions.
- Please use Python 3.5 or 3.6 (for full support of typing annotations). You have to make the functions’ return values match the required type.
- In this programming assignment you will implement k-Nearest Neighbours. We have provided the bootstrap code and you are expected to complete the classes and functions.
- Download all files of PA1 from Vocareum and save in the same folder.
- Only modifications in files {knn.py, utils.py} will be accepted and graded. test.py can be used testing purposes on your local system for your convenience. It will not be graded on vocareum. Submit {knn.py, utils.py} on Vocareum once you are finished. Please delete unnecessary files before you submit your work on Vocareum.
- DO NOT CHANGE THE OUTPUT FORMAT. DO NOT MODIFY THE CODE UNLESS WE INSTRUCT YOU TO DO SO. A homework solution that mismatches the provided setup, such as format, name initializations, etc., will not be graded. It is your responsibility to make sure that your code runs well on Vocareum.
Notes on distances and F-1 score
In this task, we will use four distance functions: (we removed the vector symbol for simplicity)
- Canberra Distance
- Minkowski Distance
- Euclidean distance
- Inner product distance
- Gaussian kernel distance
- Cosine Similarity
An inner product is a generalization of the dot product. In a vector space, it is a way to multiply vectors together, with the result of this multiplication being a scalar.
Cosine Distance = 1 - Cosine Similarity
F1-score is a important metric for binary classification, as sometimes the accuracy metric has the false positive (a good example is in MLAPP book 2.2.3.1 “Example: medical diagnosis”, Page 29). We have provided a basic definition. For more you can read 5.7.2.3 from MLAPP book.
Part 1.1 F-1 score and Distances
Implement the following items in utils.py
- function f1_score
- class Distances
- function canberra_distance
- function minkowski_distance
- function euclidean_distance
- function inner_product_distance
- function gaussian_kernel_distance
- function cosine distance
Simply follow the notes above and to finish all these functions. You are not allowed to call any packages which are already not imported. Please note that all these methods are graded individually so you can take advantage of the grading script to get partial marks for these methods instead of submitting the complete code in one shot.
Part 1.2 KNN Class
The following functions are to be implemented in knn.py:
Part 1.3 Hyperparameter Tuning
In this section, you need to implement tuning_without_scaling function of HyperparameterTuner class in utils.py. You should try different distance functions you implemented in part 1.1, and find the best k. Use k range from 1 to 30 and increment by 2. Use f1-score to compare different models.
Part 2 Data transformation
We are going to add one more step (data transformation) in the data processing part and see how it works. Sometimes, normalization plays an important role to make a machine learning model work. This link might be helpful https://en.wikipedia.org/wiki/Feature_scaling
Here, we take two different data transformation approaches.
Normalizing the feature vector
This one is simple but some times may work well. Given a feature vector, the normalized feature vector is given.
If a vector is a all-zero vector, we let the normalized vector also be a all-zero vector.
Min-max scaling the feature matrix
The above normalization is data independent, that is to say, the output of the normalization function doesn’t depend on rest of the training data. However, sometimes it is helpful to do data dependent normalization. One thing to note is that, when doing data dependent normalization, we can only use training data, as the test data is assumed to be unknown during training (at least for most classification tasks).
The min-max scaling works as follows: after min-max scaling, all values of training data’s feature vectors are in the given range. Note that this doesn’t mean the values of the validation/test data’s features are all in that range, because the validation/test data may have different distribution as the training data.
Implement the functions in the classes NormalizationScaler and MinMaxScaler in utils.py
- normalize
normalize the feature vector for each sample . For example, if the inputfeatures = [[3, 4], [1, -1], [0, 0]], the output should be[[0.6, 0.8], [0.707107, -0.707107], [0, 0]] - min_max_scale
normalize the feature vector for each sample . For example, if the inputfeatures = [[2, -1], [-1, 5], [0, 0]], the output should be[[1, 0], [0, 1], [0.333333, 0.16667]]
Hyperparameter tuning with scaling
This part is similar to Part 1.3 except that before passing your trainig and validation data to KNN model to tune k and distance function, you need to create the normalized data using these two scalers to transform your data, both training and validation. Again, we will use f1-score to compare different models. Here we have 3 hyperparameters i.e. k, distance_function and scaler.
Use of test.py file
Please make use of test.py file to debug your code and make sure your code is running properly. After you have completed all the classes and functions mentioned above, test.py file will run smoothly and will show a similar output as follows (your actual output values might vary).
Grading Guideline for KNN
- F-1 score and Distance functions
- MinMaxScaler and NormalizationScaler
- Finding best parameters before scaling
- Finding best parameters after scaling
- Doing classification of the data